Title
ProfessorOffice Building
Chicoine Architecture, Mathematics and Engineering HallOffice
258Mailing Address
Chicoine Architecture, Math & Engineering Building 258Math & Statistics-Box 2225
University Station
Brookings, SD 57007
Biography
Dr. Ge's teaching and research focus on bioinformatics, artificial intelligence (AI), and data science. Bioinformatics is a discipline that combines statistics, computer science, and biological context to analyze large genomics datasets.Education
2000 PhD in Engineering, University of Tokyo, Japan1997 MS in Condensed Matter Physics, Beijing University of Science and Technology, China
1994 BS in Applied Physics, Beijing University of Science and Technology, China
Academic Interests
Data Science, AI, Bioinformatics, GenomicsAcademic Responsibilities
STAT 435/535 Applied Bioinformatics, spring semesters for students with biological backgroundSTAT 736 Bioinformatics, fall semester for students in statistics, computer science departments or others with programming skills
STAT 414/514 Basic R programming, spring semester, one credit online course.
STAT 492: Data visualization using R.
STAT 442 Exploratory and Cloud-based Data Analysis. A project-based course where students analyze real datasets and present and discuss every week.
Grants
National Institutes of Health (NIH): An interactive tool for in-depth and reproducible analysis of RNA-seq data; Award Number:R01HG010805; Principal Investigator (PI): Xijin Ge; Organization: SDSU; Start Date: 09/2020; Award Amount: $869,378NIH: The Immune regulation of macrophage antibody dependent cellular phagocytosis, Award Number: U01AI148153; PI: Adam Hoppe; Role: Consultant; Organization: SDSU; Start Date: 07/2020; Award Amount: ~$1,500,000
NIH: BioSystems Networks and Translational Research - Insights into Inflammation (BioSNTR-II), Award Number: P20GM135008; PI: Adam Hoppe; Role: Core co-director; Start Date: 03/2022; Total award amount: ~$12,000,000
NIH: Adventures in Drug Discovery: Integrating Data Science into the Science Curriculum, Award Number: R25GM150143; PI: Charles Xie; Role: Consultant; Organization: Institute for Future Intelligence; Start Date: 07/2023; Total award amount: ~$1,000,000
National Science Foundation & the State of South Dakota: The Biochemical Spatiotemporal NeTwork Resource (BioSNTR): Building a bio-Economy in South Dakota, PI: James Rice, Start Date: 07/2013; Role: Participant; Organization: SD EPSCoR; Total award amount: ~25,000,000.
NIH: Large-scale expression analysis of natural antisense transcripts, Award number: R01GM083226, PI: Xijin Ge; Role: PI, Organization: SDSU; Start Date: 04/2009; Award amount: $835,586
Creative Activities
Selected publications:Ge X, D Jung, ShinyGO: a graphical enrichment tool for animals and plants, Bioinformatics, 36:2628-2629, 2020.
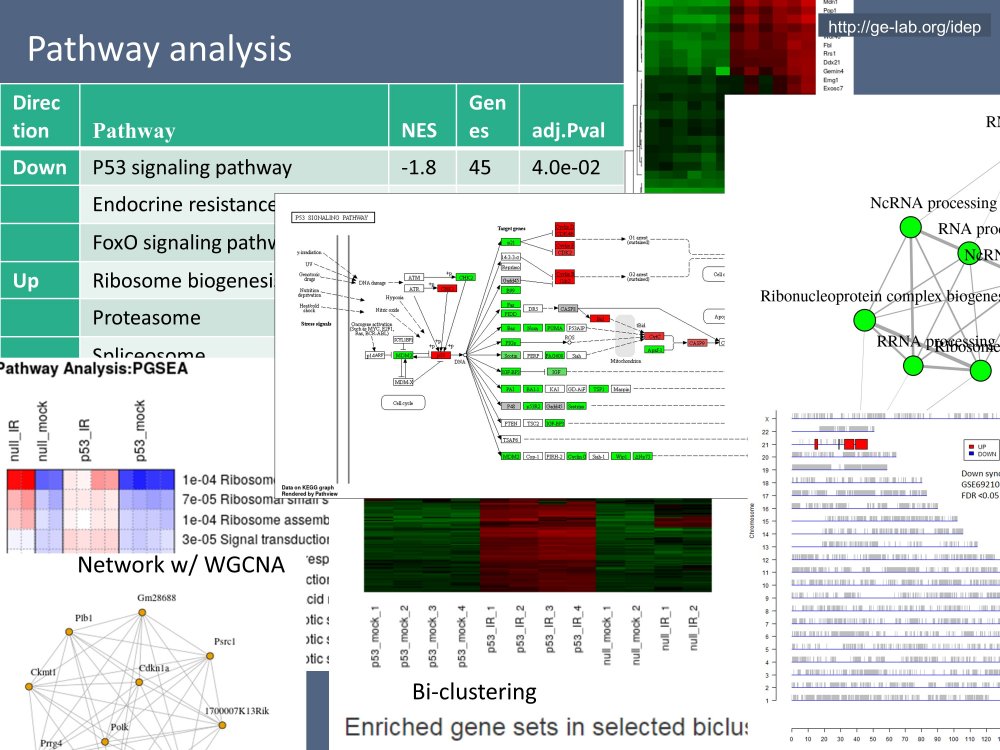
Ge S.X., Son EW, Yao RN, iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data, BMC Bioinformatics 19:124, 2018.
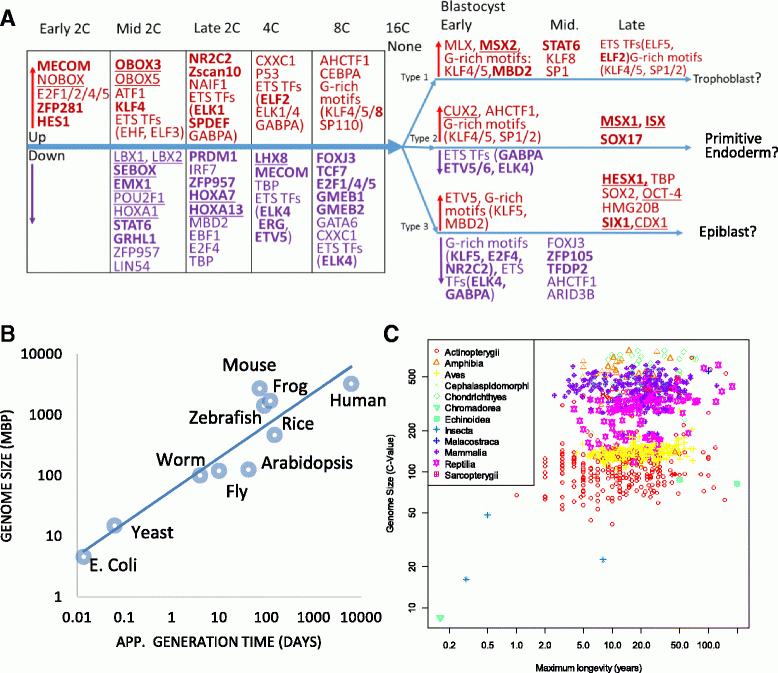
Ge SX. Exploratory bioinformatics investigation reveals importance of "junk" DNA in early embryo development. BMC Genomics. 18(1):200, 2017.
Ling MH, Ban Y, Wen H, Wang SM, Ge SX. Conserved expression of natural antisense transcripts in mammals. BMC Genomics. 14:243, 2013.
Shi L, et al. The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models. Nature Biotechnology, 28(8):827-38, 2010.
Ge X, Iwata S. Learning the parts of objects by auto-association. Neural Network.15(2):285-95, 2002.
Free online Book:
Learn R through Examples, https://gexijin.github.io/learnR/
Area(s) of Research
Currently, the main focus of his group is the development of tools (iDEP & ShinyGO) and resources (pathway databases) for the interpretation of large genomic data. Another emphasis is using AI, especially Large Language Models like ChatGPT, in helping analyzing and visualizing data.Applications of Research
Bioinformatics tools (iDEP, ShinyGO) developed by Dr Ge's group have been used hundreds of thousands of times by scientists across the world to study various processes ranging from agriculturally-important crops to human diseases such as cancer.Department(s)
Image for Department of Mathematics and Statistics
Department of Mathematics and Statistics
Links
Dr. Ge's publicationsGe lab softwareDr Ge on LinkedInDr. Ge on TwitterOpen Prairie
Open PRAIRIE - Public Research Access Institutional Repository and Information Exchange