Title
Associate Professor and Winter Wheat BreederOffice Building
Young Brothers Seed Technology LaboratoryOffice
113BMailing Address
Seed Technology Lab 113BAgronomy, Horticulture & Plant Science-Box 2108
University Station
Brookings, SD 57007
Biography
Dr. Sehgal received his Ph.D. in Plant Breeding and Genetics from Punjab Agricultural University. He worked as a research scientist at Kansas State University (2006-2014). Since joining SDSU in 2014 he is leading the winter wheat breeding program at SDSU. His research is focused on developing and releasing high-yield winter wheat cultivars with resistance to biotic (mostly FHB, stripe, leaf, and stem rust, and WSMV) and abiotic stresses (winter hardiness and drought). In addition to research, Dr. Sehgal has also been active in teaching, student advising, and outreach for the wheat industry in South Dakota. Dr. Sehgal has published more than 70 peer-reviewed articles including Nature, Science, PNAS, Genome Biology, Plant Physiology, New Phytologist, Genetics, and Theoretical and Applied Genetics.Education
Ph.D. Plant BreedingM.S. Plant Breeding
B. S. Agriculture (Honors Plant Breeding & Seed Technology)
Academic Interests
Wheat Breeding and GeneticsExpertise
Academic Responsibilities
PS787 -Advanced Plant Breeding (Spring odd year)Committee Activities
National Plant Breeding Coordinating CommitteeUSWBSI-HWW-CP (Chair)
National Wheat Improvement Committee
SDSU Variety Release Committee
Extension Responsibilities
No assigned responsibilities, but I make presentations in more than 6 field days on producer farms and SDSU research stations across the state educating the growers about the winter wheat varieties available and their characteristics.Awards and Honors
2023 'F.O.Bulter Award Excellence in Research' at South Dakota State University, Brookings, SD2023 ‘AHPS Excellence in Research award in the SDSU AHPS department, Brookings, SD
2023 'Best of Show’ Millers award (2022) for best quality wheat varieties, Wheat Quality Council (WQC), Kansas City, KS
2022 ‘AHPS Excellence in Research award in the SDSU AHPS department, Brookings, SD
2022 'Best of Show’ Millers award (2021) for best quality wheat varieties, Wheat Quality Council (WQC), Kansas City, MO
2010 "Early Career Speaker Award", International Wheat Genome Sequencing Consortium.
2010 “K-State Make A Difference Award” for undergraduate mentoring, Kansas State University, Manhattan KS.
2004 "University Randhawa Gold Medal" for the best essay on Second Green Revolution, Punjab Agricultural University, India.
2003 University Merit Scholarship (Ph.D. program), Punjab Agricultural University, India.
Grants
Grants Received as PICurrent Grants
1. Developing high yielding winter wheat varieties with excellent end-use quality for South Dakota funded by South Dakota Wheat Commission from 2023 to 2024.
2. Accelerated breeding for disease and insect pest resistance in winter wheat funded by South Dakota Wheat Commission from 2023 to 2024.
3. Winter Wheat Breeding for Scab Resistance in South Dakota funded by USDA-USWBSI from 2022-2025.
4. WheatCAP: WheatCAP: Innovation in Genomic Technology to Accelerate Breeding: “Leveraging high-throughput genotyping and phenotyping technologies to accelerate wheat improvement (2022-2026)
5. Breeding for improved nitrogen use efficiency (NUE) in South Dakota winter wheat under regenerative agriculture management (SD NREC, 2022-2024)
Patents
Plant Variety Protection (PVP)'Oahe' 2019
'Thompson' 2019
'Winner' 2021
'Draper' 2021
'SD Andes' 2022
'SD Midland' 2023
Professional Memberships
Crop Science Society of AmericaAgronomy Society of America
American Association for the Advancement of Science
Coordinating committee member International Wheat Genome Sequencing Consortium (IWGSC) (2009-2014)
Work Experience
Associate Professor and Winter Wheat Breeder, South Dakota State UniversityAssistant Professor and Winter Wheat Breeder, South Dakota State University, 2014-2020
Senior Scientist, Kansas State University, Manhattan, KS from Jul 2013 - Sep 2014
Research Associate, Kansas State University, Manhattan, KS from Nov 2006 - Jun2013
Area(s) of Research
The focus of SDSU-Hard Winter Wheat (HWW) breeding program is a continual genetic enhancement of HWW productivity (both quantity and quality) and value addition for growers in South Dakota and the Northern Great Plains.Winter Wheat Varieties Released:
1. Hard red winter (HRW) wheat cultivar ‘Oahe’ was released by SDAES (8/2016). Oahe offers a combination of high yield with good test weight and moderate resistance to stripe rust, leaf rust, wheat streak mosaic virus, and Fusarium head blight.
2. Hard red winter (HRW) wheat cultivar ‘Thompson’ was released by SDAES (11/2017). Thompson is a high yielding taller semi-dwarf HRW variety adapted to central South Dakota with moderate resistance to leaf rust and stem rust and acceptable milling quality.
3. Hard red winter (HRW) wheat cultivar ‘Winner’ was released by SDAES (9/2019). Winner is semi-dwarf variety with unusually broad adaptation to the eastern half of the Northern Great Plains. Winner has medium height and medium maturity with higher yield potential, good baking quality, and moderate resistance to stem rust.
4. Hard red winter (HRW) wheat cultivar ‘Draper’ was released by SDAES (12/2019). Draper is a semi-dwarf variety with good straw strength and a limited target region, specifically western South Dakota. Draper has improved yield potential with average test weight, grain protein, and acceptable milling and baking quality. It is resistant to soilborne mosaic virus.
5. Hard red winter (HRW) wheat cultivar ‘SD ANDES’ (12/2020). SD Andes is a semi-dwarf variety with late maturity. It has excellent straw strength, winter hardiness. SD Andes has good resistance to stripe rust and average tolerance to FHB. SD Andes has improved yield potential with average test weight, grain protein, and acceptable milling and baking quality. It is best adapted to eastern South Dakota and the neighboring region.
6. Hard red winter (HRW) wheat cultivar ‘SD MIDLAND’ (12/2021). SD Midland is a semi-dwarf variety with medium-tall in height and late in maturity. It is high-yielding with average protein and test weight and excellent milling and baking quality. SD Midland is moderately resistant to stripe rust and yields well in lower rainfall areas. It is a bronze chaff winter wheat and would be a good replacement for later maturing varieties like Redfield.
7. Hard red winter (HRW) wheat cultivar ‘SD Pheasant’ released in Fall 2023. SD Pheasant is a high-yielding line with good test weight and grain protein content. SD Pheasant is resistant to leaf rust and moderately resistant to stem rust, hessian fly and moderately tolerant to stripe rust, FHB, and Bacterial Leaf Streak. Along with excellent grain yield potential, SD Pheasant has good milling characteristics and excellent baking characteristics. It was awarded millers choice ‘Best of Show’ award in 2023 for excellent milling and baking characteristics.
Publications: Refereed Journal Articles
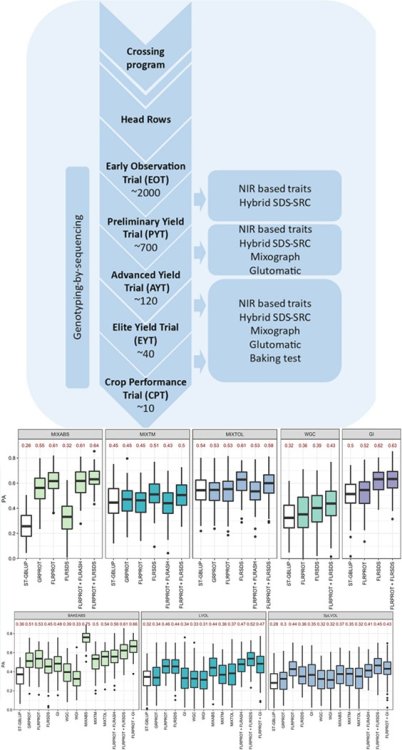
70. Gill, H.S., Brar, N., Halder, J., Hall, C., Seabourn, B.W., Chen, Y.R., St. Amand, P., Bernardo, A., Bai, G., Glover, K. and Turnipseed, B. and Sehgal, S.K.*, 2023. Multi‐trait genomic selection improves the prediction accuracy of end‐use quality traits in hard winter wheat. The Plant Genome, p.e20331. https://doi.org/10.1002/tpg2.20331
69. Kaur, S., Gill, H.S., Breiland, M., Kolmer, J.A., Gupta, R., Sehgal, S.K.*, and Gill, U., 2023. Identification of leaf rust resistance loci in a geographically diverse panel of wheat using genome-wide association analysis. Frontiers in Plant Science, 14, p.1090163.
68. Ma, C., Qian, J., Feng, Y., Sehgal, S.K., Zhao, Y., Chen, Q., Li, H. and Liu, W., 2023. Genetic Mapping of a Novel Gene PmAege7M from Aegilops geniculata Conferring Resistance to Wheat Powdery Mildew. Plant Disease, https://doi.org/10.1094/PDIS-04-23-0764-RE
67. Lopez, S.R., Wiersma, A.T., Strauss, N.M., Watkins, T., Baik, B.K., Zhang, G., Sehgal, S.K., Kolb, F.L., Poland, J.A., Mason, R.E. and Carter, A.H., 2023. Description of U6719‐004 wheat germplasm with YrAS2388R stripe rust resistance introgression from Aegilops tauschii. Journal of Plant Registrations, 17(1), pp.26-33
66. Liu, Z., Li, Y., Bernardo, A., Amand, P.S., Zhang, P., Sehgal, S. and Bai, G., 2023. Development of diagnostic SNP markers for identification of rye 1RS translocations in wheat. Crop Science, 63(1), pp.255-265
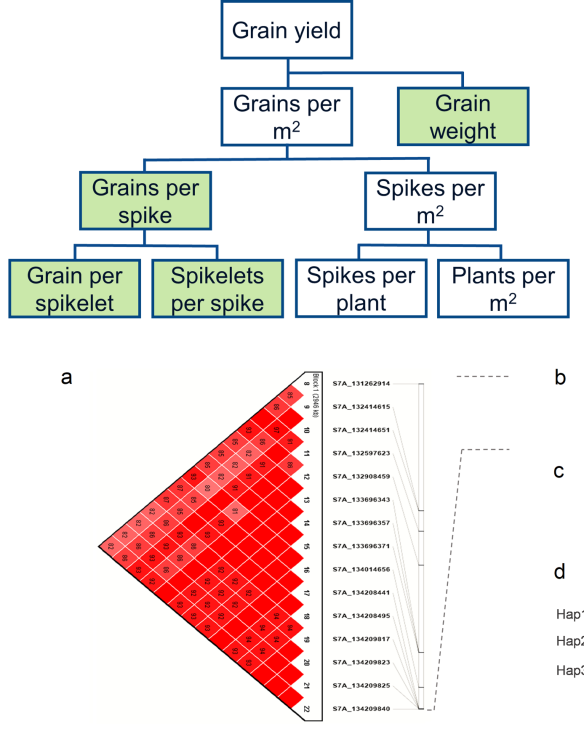
65. Halder, J., Gill, H.S., Zhang, J., Altameemi, R., Olson, E., Turnipseed, B. and Sehgal, S.K.*, 2023. Genome‐wide association analysis of spike and kernel traits in the US hard winter wheat. The Plant Genome, 16(1), p.e20300. https://doi.org/10.1002/tpg2.20300
64. Gill, H.S., Halder, J., Zhang, J., Rana, A., Kleinjan, J., Amand, P.S., Bernardo, A., Bai, G. and Sehgal, S.K.,* (2022) Whole-genome analysis of hard winter wheat germplasm identifies genomic regions associated with spike and kernel traits. Theoretical and Applied Genetics, 135(9), pp.2953-2967.
63. Zhang, J., Gill, H.S., Halder, J., Brar, N.K., Ali, S., Bernardo, A., Amand, P.S., Bai, G., Turnipseed, B. and Sehgal, S.K.,* (2022) Multi-locus genome-wide association studies to characterize Fusarium head blight (FHB) resistance in hard winter wheat. Frontiers in Plant Science, 13: 946700.
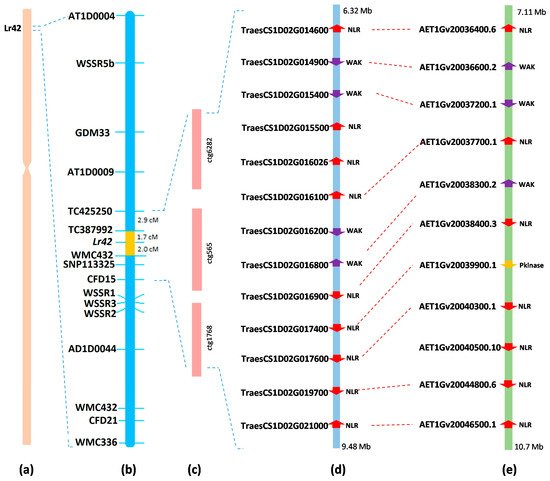
62. Lin, G., Chen, H., Tian, B., Sehgal, S.K., Singh, L., Xie, J., Rawat, N., Juliana, P., Singh, N., Shrestha, S. and Wilson, D.L. (2022) Cloning of the broadly effective wheat leaf rust resistance gene Lr42 transferred from Aegilops tauschii. Nature Communications, 13(1), pp.1-12.
61. He F, Wang W, Rutter WB, Jordan KW, Ren J, Taagen E, DeWitt N, Sehgal D, Sukumaran S, Dreisigacker S, Reynolds M, Halder J, Sehgal SK, Liu S, Akhunov E (2022). Genomic variants affecting homoeologous gene expression dosage contribute to agronomic trait variation in allopolyploid wheat. Nat. Commun 13, 826
60. Zhang, J., Gill, H.S., Brar, N.K., Halder, J., Ali, S., Liu, X., Bernardo, A., Amand, P.S., Bai, G., Gill, U.S. and Turnipseed, B. Sehgal, S.K.*, 2022. Genomic prediction of Fusarium head blight resistance in early stages using advanced breeding lines in hard winter wheat. The Crop Journal 10:1675-104
59. Liu, Z., Li, Y., Bernardo, A., Amand, P.S., Zhang, P., Sehgal, S., Bai, G., (2022). Development of diagnostic SNP markers for identification of rye 1RS translocations in wheat translocations in wheat. Crop Sci (Early view). https://doi.org/10.1002/csc2.20841
58. Men, W., Fan, Z., Ma, C., Zhao, Y., Wang, C., Tian, X., Chen, Q., Miao, J., He, J., Qian, J. and Sehgal, S.K. Li, H, Liu .W. (2022) Mapping of the novel powdery mildew resistance gene Pm2Mb from Aegilops biuncialis based on ph1b-induced homoeologous recombination. Theoretical and Applied Genetics 135:2993-3003.
57. Tian, X., Chen, Q., Ma, C., Men, W., Liu, Q., Zhao, Y., Qian, J., Fan, Z., Miao, J., He, J., Sehgal, S.K., Li, H, Liu .W. (2022) Development and characterization of Triticum aestivum-Aegilops longissima 6Sl recombinants harboring a novel powdery mildew resistance gene Pm6Sl. Frontiers in Plant Science, 13:918508.
56. Lopez S, Wiersma A, Strauss N, Watkins T, Baik B, Zhang G, Sehgal S, Kolb FL, Poland J, Mason E, Carter AH, Olson E (2022) Description of U6719-004 wheat germplasm with YrAS2388R stripe rust resistance introgression from Aegilops tauschii. J Plant Registrations 2022;1–8
55. Gill, H.S., Halder, J., Zhang., J, Brar, N., Rai, T.S., Hall, C., Bernardo, A., St Amand, P., Bai, G., Olson, E., Ali, S., Turnipseed, B., Sehgal, S.K.,* (2021) Multi-trait multi-environment genomic prediction of agronomic traits in advanced breeding lines of winter wheat. Front. Plant Sci., 18 August 2021 | https://doi.org/10.3389/fpls.2021.709545
54. AlTameemi, R., Gill, H.S., Ali, S.... Sehgal (2021) Genome-wide association analysis permits characterization of Stagonospora nodorum blotch (SNB) resistance in hard winter wheat. Sci Rep 11, 12570 (2021). https://doi.org/10.1038/s41598-021-91515-6
53. Li H, X Tian, S Pei, W Men, C Ma, SK Sehgal, Y Zhao, Q Chen, B Wang, et al (2021) Development of Novel Wheat-Aegilops longissima 3Sl Translocations Conferring Powdery Mildew Resistance and Specific Molecular Markers for Chromosome 3Sl Plant Disease
52. Liu S, Wang D, Lin M, Sehgal SK, Dong L, Wu Y, Bai G (2020) Artificial selection in breeding extensively enriched a functional allelic variation in TaPHS1 for pre-harvest sprouting resistance in wheat. Theor Appl Genet (in press)
51. Sidhu JS, Singh D, Gill HS, Brar NK, Qiu Y, Halder J, Al Tameemi R, Turnipseed B, Sehgal SK* (2020) Genome-wide association study uncovers novel genomic regions associated with coleoptile length in hard winter wheat. Front. Genet. 10:1345
50. Latif S,Wang L, Khan J, Ali Z, Sehgal SK, Babar MA, Wang J, Quraishi UM (2020) Deciphering the role of Stay-Green Trait to mitigate terminal heat stress in Bread Wheat. Agronomy 10(7): 1001
49. Kaur N, Sehgal SK, Glover D, Byamukama E, Ali S (2020) Impact of Fusarium graminearum on seed germination and seedling blight in hard red spring cultivars in South Dakota. J Plant Pathology & Microbiology 11:495
48. Liu W, Li H, Dong Z, Ma C, Xia Q, Tian X, Sehgal S, Friebe B, Ma P (2020) A spontaneous wheat-Aegilops longissima translocation carrying Pm66 confers resistance to powdery mildew. Theor Appl Genet 133(4):1149-1159
47. Dong Z, X, Ma C, Xia Q, Wang B, Chen Q, Sehgal SK, Friebe B, Li H and Liu W (2020) Towards Physical Mapping of Pm57, a Powdery Mildew Resistance Gene Derived from Aegilops searsii. Int. J. Mol. Sci. 21(1), 322; https://doi.org/10.3390/ijms21010322
46. Halder J, Zhang J, Ali S, Sidhu JS, Gill HS, Talukdar S, Kleinjan J, Turnipseed B, Sehgal SK* (2019) Mining and genomic characterization of resistance to tan spot, Stagonospora nodorum blotch (SNB), and Fusarium head blight in Watkins core collection of wheat landraces. BMC Plant Biology 19:480
45. Ramakrishnan SM, Sidhu JS, Ali S, Kaur N, Wu J, Sehgal SK* (2019) Molecular characterization of bacterial leaf streak resistance in hard winter wheat. PeerJ 7:e7276
44. Gill HS, Li C, Sidhu JS, Liu W, Wilson D, Bai G, Gill BS, Sehgal SK* (2019) Fine mapping of the wheat leaf rust resistance gene. Int. J. Mol. Sci. 20:2445; https://doi.org/10.3390/ijms20102445
43. Sidhu JS, Ramakrishnan SM, Ali S, Bernardo A, Bai G, Abdullah S, Ayana G, Sehgal SK*. (2019) Assessing the genetic diversity and characterizing genomic regions conferring tan spot resistance in cultivated rye. PLoS ONE 14: e0214519.
42. Djanaguiraman M, Prasad PVV, Kumari J, Sehgal SK, Friebe B, Dalovic I, Chen Y, Siddique KHM, Gill BS (2019) Alien chromosome segment from Aegilops speltoides and Dasypyrum villosum increases drought tolerance in wheat via profuse and deep root system. BMC Plant Biology 19:242.
41. Singh N, Wu S, Raupp J, Sehgal S, Arora S, Tiwari V, Vikram P, Singh S, Chhuneja P, Gill BS, Poland J (2019) Efficient curation of genebanks using next generation sequencing reveals substantial duplication of germplasm accessions. Scientific Reports 9: 650.
40. Singh N, Wu S, Tiwari V, Sehgal S, Raupp J, Wilson D, Abbasov M, Gill BS, Poland J (2019) Genomic analysis confirms population structure and identifies inter-lineage hybrids in Aegilops tauschii. Front. Plant Sci. 10:9.
39. Kumar S, Sieverding H, Lai L, Thandiwe N, Wienhold B, Redfearn D, Archer D, Ussiri D, Faust D, Landblom D, Grings E, Stone JJ, Jacquet J, Pokharel K, Liebig M, Schmer M, Sexton P, Mitchell R, Smalley S, Osborne S, Ali S, Şentürklü S, Sehgal S, Owens V, Jinc V (2019) Facilitating crop-livestock reintegration in the Northern Great Plains. Agronomy J 111:1–16 (Peer-reviewed review article)
38. Appels R, Eversole K, Feuille, C, Keller B, Rogers J, Stein N, Pozniak CJ, Choulet F, Distelfeld A, Poland J, Ronen G, Sharpe AG, Pozniak C, Barad O, Baruch K, Keeble-Gagnère G, Mascher M, Ben-Zvi G, Josselin A-A, Himmelbach A, Balfourier F, Gutierrez-Gonzalez J, Hayden M, Koh C, Muehlbauer G, Pasam RK, Paux E, Rigault P, Tibbits J, Tiwari V, Spannagl M, Lang D, Gundlach H, Haberer G, Mayer KFX, Ormanbekova D, Prade V, Šimková H, Wicker T, Swarbreck D, Rimbert H, Felder M, Guilhot N, Kaithakottil G, Keilwagen J, Leroy P, Lux T, Twardziok S, Venturini L, Juhász A, Abrouk M, Fischer I, Uauy C, Borrill P, Ramirez-Gonzalez RH, Arnaud D, Chalabi S, Chalhoub B, Cory A, Datla R, Davey MW, Jacobs J, Robinson SJ, Steuernagel B, van Ex F, Wulff BBH, Benhamed M, Bendahmane A, Concia L, Latrasse D, Alaux M, Bartoš J, Bellec A, Berges H, Doležel J, Frenkel Z, Gill B, Korol A, Letellier T, Olsen O-A, Singh K, Valárik M, van der Vossen E, Vautrin S, Weining S, Fahima T, Glikson V, Raats D, Číhalíková J, Toegelová H, Vrána J, Sourdille P, Darrier B, Barabaschi D, Cattivelli L, Hernandez P, Galvez S, Budak H, Jones JDG, Witek K, Yu G, Small I, Melonek J, Zhou R, Belova T, Kanyuka K, King R, Nilsen K, Walkowiak S, Cuthbert R, Knox R, Wiebe K, Xiang D, Rohde A, Golds T, Čížková J, Akpinar BA, Biyiklioglu S, Gao L, N’Daiye A, Kubaláková M, Šafář J, Alfama F, Adam-Blondon A-F, Flores R, Guerche C, Loaec M, Quesneville H, Condie J, Ens J, Maclachlan R, Tan Y, Alberti A, Aury J-M, Barbe V, Couloux A, Cruaud C, Labadie K, Mangenot S, Wincker P, Kaur G, Luo M, Sehgal S, Chhuneja P, Gupta OP, Jindal S, Kaur P, Malik P, Sharma P, Yadav B, Singh NK, Khurana J, Chaudhary C, Khurana P, Kumar V, Mahato A, Mathur S, Sevanthi A, Sharma N, Tomar RS, Holušová K, Plíhal O, Clark MD, Heavens D, Kettleborough G, Wright J, Balcárková B, Hu Y, Salina E, Ravin N, Skryabin K, Beletsky A, Kadnikov V, Mardanov A, Nesterov M, Rakitin A, Sergeeva E, Handa H, Kanamori H, Katagiri S, Kobayashi F, Nasuda S, Tanaka T, Wu J, Cattonaro F, Jiumeng M, Kugler K, Pfeifer M, Sandve S, Xun X, Zhan B, Batley J, Bayer PE, Edwards D, Hayashi S, Tulpová Z, Visendi P, Cui L, Du X, Feng K, Nie X, Tong W, Wang L (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science. 361:eaar7191.
37. Ayana GT, Ali S, Sidhu JS, Gonzalez Hernandez JL, Turnipseed B, Sehgal SK* (2018) Genome-wide association study for spot blotch resistance in hard winter wheat. Front. Plant Sci. 9:926.
36. Wiersma AT, Whetten RB, Zhang G, Sehgal SK, Kolb FL, Poland JA, Mason RE, Carter AH, Cowger C, Olson EL (2018) Registration of two wheat germplasm lines fixed for Pm58. J. Plant. Reg. 12: 270-273.
35. Abdullah S, Sehgal SK, Glover K, Ali S (2017) Reaction of global collection of rye (Secale cereale L.) to tan spot and Pyrenophora tritici-repentis races in South Dakota. The Plant Pathology Journal 33: 229-237.
34. Abdullah S, Sehgal SK, Ali S (2017) Race diversity of Pyrenophora tritici-repentis in South Dakota and response of predominant wheat cultivars to tan spot. J Plant Pathol Microbiol 8: 409.
33. Abdullah S, Sehgal SK*, Yu J, Turnipseed B, Ali S* (2017) Insight into tan spot and stem rust resistance and susceptibility by studying the pre-green revolution global collection of wheat. The Plant Pathology Journal 33:125-132.
32. Abdullah S, Sehgal SK, Ali S, Laiiatukas Ž, Ittu M, Kaur N (2017) Characterization of Pyrenophora tritici-repentis (tan spot of wheat) races in Baltic States and Romania. The Plant Pathology Journal: 33(2): 133–139.
31. Gong W, Gong W, Han R, Li G, Sehgal SK, Li H, Liu H, Song J, Song G, Liu C, Liu J (2016) Chromosome arm-specific markers from Aegilops searsii permits targeted introgression. Biologia 71:87-92.
30. Wiersma AT, Brown LK, Brisco EI, Liu TL, Childs KL, Poland JA, Sehgal SK, Olson EL (2016) Fine mapping of the stem rust resistance gene SrTA10187. Theor Appl Genet 128: 2369-2378.
29. Davis MA, Bockus W, Zhang G, Fritz AK, Baenziger PS, Marais GF, Sehgal SK (2016) Reaction of Kansas, Nebraska, South Dakota, and North Dakota winter wheat accessions to Fusarium head blight (FHB). Plant Disease Management Reports Vol 11:CF037.
28. Chapman JA, Mascher M, Buluç AN, Barry K, Georganas E, Session A, Strnadova V, Jenkins J, Sehgal S, Oliker L, Schmutz J, Yelick KA, Scholz U, Waugh R, Poland JA, Muehlbauer GJ, Stein N, Rokhsar DS (2015) A whole-genome shotgun approach for assembling and anchoring the hexaploid bread wheat genome. Genome Biol. 16:26.
27. Bockus W, Zhang G, Fritz AK, Davis MA, Baenziger PS, Marais GF, Sehgal SK (2015) Reaction of Kansas, Nebraska, South Dakota, and North Dakota winter wheat accessions to Fusarium head blight (FHB). Plant Disease Management Reports Vol 10: CF043.
26. Liu S, Sehgal SK, Lin M, Li J, Trick H, Gill BS, Bai G (2015) Independent mis-splicing mutations in TaPHS1 causing loss of pre-harvest sprouting (PHS) resistance during wheat domestication. New Phytologist 208:928-35.
25. Chhuneja P, Yadav B, Stirnweis D, Hurni S, Kaur S, Elkot AF, Keller B, Wicker T, Sehgal SK, Gill BS, Singh KS (2015) Fine mapping of a new powdery mildew resistance gene PmTb7A and a new allele of Pm1Tb in Triticum boeoticum and identification of STS markers suited for MAS using the survey sequence data of chromosome 7AL. Theor Appl Genet 128:2099-2111.
24. Cainong JC, Bockus WW, Feng Y, Chen P, Qi L, Sehgal SK, Danilova TV, Koo D, Friebe B, Gill BS (2015) Chromosome engineering, mapping, and transfer of native grass resistance to Fusarium Head Blight disease into wheat. Theor Appl Genet 128:1019-1027.
23. Koo D-H, Sehgal SK, Friebe B, Gill BS (2015) Structure and stability of telocentric chromosomes in wheat. PLoS ONE 10(9): e0137747.
22. Jugulam M, Niehues K, Godar AS, Koo DH, Danilova TV, Friebe BR, Sehgal SK, Varanasi V, Wiersma A, Westra P, Stahlman P, Gill BS (2014) Tandem amplification of a chromosomal segment harboring EPSPS locus confers glyphosate resistance in Kochia scoparia. Plant Physiol.166: 1200-1207.
21. Tiwari VK, Wang S, Sehgal S, Vrána J, Friebe B, Kubaláková M, Chhuneja P, Doležel J, Akhunov E, Kalia B, Sabir J, Gill BS (2014) SNP discovery and mapping alien introgressions in wheat. BMC Genomics 15: 274.
20. Gawroński P, Ariyadasa R, Poursarebani N, Himmelbach A, Kilian B, Stein N, Steuernagel B, Hensel G, Kumlehn J, Sehgal SK, Gill BS, Gould P, Hall A, Schnurbusch T (2014) A distorted circadian clock causes early flowering and temperature-dependent variation in spike development in the eps-3am mutant of einkorn wheat. Genetics 196:1253-1261.
19. Liu S, Sehgal SK, Li J, Lin M, Trick H, Yu J, Gill BS, Bai G (2013) Cloning and characterization of a critical regulator for pre-harvest sprouting in wheat. Genetics 195:263-273.
18. Luo M-C, Gu Y, You FM, Deal KD, MaY, Hu Y, Huo N, Wang Y, Wang J, Chen S, Jorgensen CM, McGuire PE, Stein J, Ware D, McCombie R, Kianian SF, Martis MM, Mayer K, Sehgal SK, Gill BS, Bevan M, Dolezel J, Anderson OD, Dvorak J (2013) Physical map and gene space sequence of Aegilops tauschii, the source of wheat D genome, provide insights into the evolution of grass genomes. Proc. Natl. Acad. Sci. 110: 7940-7945.
17. Akhunov E*, Sehgal S*, Liang H, Wang S, Akhunova A, Kaur G, Li W, Forrest KL, See D, Simkova H, Hayden M, Luo M-C, Farris J, Dolezel J, Gill BS (2013) Comparative analysis of orthologous genes in grass genomes reveals the accelerated rates of alternative splicing and coding sequence evolution in polyploid wheat. Plant Physiol 161:252–265.
16. Brenchley R, Spannagl M, Pfeifer M, Barker GLA, D’Amore R, Allen AM, McKenzie N, Kramer K, Kerhornou A, Bolser D, Kay S, Waite D, Trick M, Bancroft I, Gu Y, Huo N, Luo M-C, Sehgal S, Gill B, Kianian K, Anderson O, Kersey P, Dvorak J, McCombie WR, Hall A, Mayer KFX, Edwards KJ, Bevan MW, Hall N (2012) Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491:705-710.
15. Sehgal SK, Li W, Rabinowicz PD, Chan A, Šimková H, Doležel J, Gill BS (2012) Chromosome arm-specific BAC end sequences permit comparative analysis of homoeologous chromosomes and genome of polyploid wheat. BMC Plant Biology 12:64.
14. Rawat N, Sehgal SK, Joshi A, Rothe N, Wilson DL, McGraw N, Vadlani PV, Li W, Gill BS (2012) A diploid wheat TILLING resource for wheat functional genomics. BMC Plant Biology 12:205.
13. Kumar S, Sehgal SK, Joshi AK, Kumar U, Prasad PVV and Gill BS (2012) Genomic characterization of drought tolerance related traits in spring wheat. Euphytica 158:265-76.
12. Liu W*, Thummasuwan S*, Sehgal SK, Chouvarine P, Peterson DG (2011) Characterization of the genome of bald cypress. BMC Genomics 12:553.
11. Sehgal SK, Gupta S, Kaur S, Sharma A, Bains NS (2011) A direct hybridization approach for gene transfer from Aegilops tauschii Coss. to Triticum aestivum L. Plant Breeding 130:98-100.
10. Gupta S, Kaur S, Sehgal S, Sharma A, Chhuneja P, Bains NS (2010) Genotypic variation for cellular thermotolerance in Aegilops tauschii Coss., the D genome progenitor of wheat. Euphytica 175: 373-381.
9. Liu C*, Li G-R*, Sehgal SK, Jia J-Q, Yang Z-J, Friebe B, Gill B (2010) Genomic divergence in the genus Dasypyrum: evidence from molecular phylogenetic analysis and in situ hybridization. Plant Systematics and Evolution 288:149–156.
8. Sharma S, Sreenivasulu N, Harshavardhan VT, Seiler C, Sharma S, Khalil ZN, Akhunov E, Sehgal SK, Röder MS (2010) Delineating the structural, functional and evolutionary relationships of sucrose phosphate synthase gene family II in wheat and related grasses. BMC Plant Biology 10:134.
7. Sehgal SK, Kaur G, Sharma I, Bains NS (2008) Development and molecular marker analysis of Karnal bunt resistant near isogenic lines in bread wheat variety PBW 343. Indian J Genet Plant Breed 68: 21-25.
6. Singh S, Sharma I, Sehgal SK, Bains NS, Guo Z, Nelson J, Bowden R (2007) Molecular mapping of QTLs for Karnal bunt resistance in two recombinant inbred populations of bread wheat. Theor Appl Genet 116: 147-154.
5. Sharma I, Bains NS, Sirari A, Sangha KS, Sehgal SK, Bodhraj (2007) Genetics of leaf and stripe rust resistance in a multiple disease resistant stock of bread wheat. Crop Improv 34:8-11.
4. Sehgal SK (2006) Studies on incorporation of Karnal bunt resistance and productivity traits from Aegilops tauschii Coss. into wheat (Triticum aestivum L.) Punjab Agricultural University, India. Doctoral Dissertation
3. Sehgal SK, Kaur G, Gupta ML (2004) Induction of androgenesis and plant regeneration in Indian mustard (Brassica juncea L. Coss.) J Pl Sci Res 20: 125-27.
2. Sehgal SK (2001) Haploid induction using anther and microspore culture in oilseed Brassicas. Punjab Agricultural University, India. Master’s Dissertation
1. Sharma S, Bajaj RK, Kaur N, Sehgal SK (2003) Combining ability studies in sunflower (Helianthus annuus L.) Crop Improv 30: 69-73.
Applications of Research
The major objectives of the breeding program are:1) Develop and release new superior winter wheat varieties with high yield stability, durable disease resistance, and improved end-use quality for South Dakota and Northern Great Plains.
2) Identify and genetically characterize germplasm with yield potential and quality traits, resistance to diseases and insect pests, drought, heat and winter hardiness and utilize them to enhance the productivity and resistance levels in winter wheat germplasm, thus leading to the development of improved varieties.
Department(s)
Image for Department of Agronomy, Horticulture and Plant Science
Department of Agronomy, Horticulture and Plant Science